CRISPR: How a Bacterial Immune Trick Became the Most Powerful Tool in Medicine
The biology behind the first approved CRISPR therapy — and what comes next
There is a moment in every scientific revolution when a new tool arrives and the entire landscape of what is possible shifts overnight. The microscope did it for cell biology. PCR did it for genetics. In 2012, a paper published in Science did it for medicine.
The tool was CRISPR-Cas9. The authors were Jennifer Doudna at UC Berkeley and Emmanuelle Charpentier, then at Umeå University in Sweden. Eight years later, they shared the Nobel Prize in Chemistry. What they had described — with elegant simplicity — was a molecular system capable of finding any sequence in a three-billion-letter genome and cutting it with surgical precision.
That sentence alone should give you pause. The human genome contains roughly three billion base pairs. CRISPR-Cas9 can find a specific twenty-letter sequence within that genome and cut it — and it can be reprogrammed to hit a different target simply by swapping out a short RNA molecule that any molecular biology lab can synthesize in days.
To understand why that is extraordinary, you need to know what came before.
Before CRISPR: A Long Road to Precision
Scientists have wanted to edit genomes since they first understood what genes were. The first tools that made it possible — called zinc finger nucleases, or ZFNs — arrived in the 1990s. They worked. But designing a ZFN for a new target required months of painstaking protein engineering and a level of expertise that kept the technology confined to a handful of specialized labs.
TALENs, the next generation, were more flexible and somewhat easier to work with. They too saw use in early gene therapy trials. But both technologies shared the same fundamental limitation: every new target required building an entirely new protein from scratch. Slow. Expensive. Inaccessible to most of the scientific world.
CRISPR-Cas9 shattered that barrier.
How It Actually Works
The name CRISPR — Clustered Regularly Interspaced Short Palindromic Repeats — describes something found in bacteria, which is where this story begins.
Bacteria have immune systems. When a virus infects a bacterium and the bacterium survives, it stores a small fragment of the viral DNA in a specific region of its own genome: the CRISPR locus. This is molecular memory. The next time the same virus appears, the bacterium transcribes that stored fragment into an RNA molecule, which acts as a recognition signal. An enzyme called Cas9 uses that RNA to find the matching viral DNA sequence — and cuts it in two, destroying the threat.
Doudna, Charpentier, and their collaborators recognised something remarkable: this system is essentially programmable. The RNA guide can be redesigned to match any DNA sequence. And Cas9 will follow it there and cut.
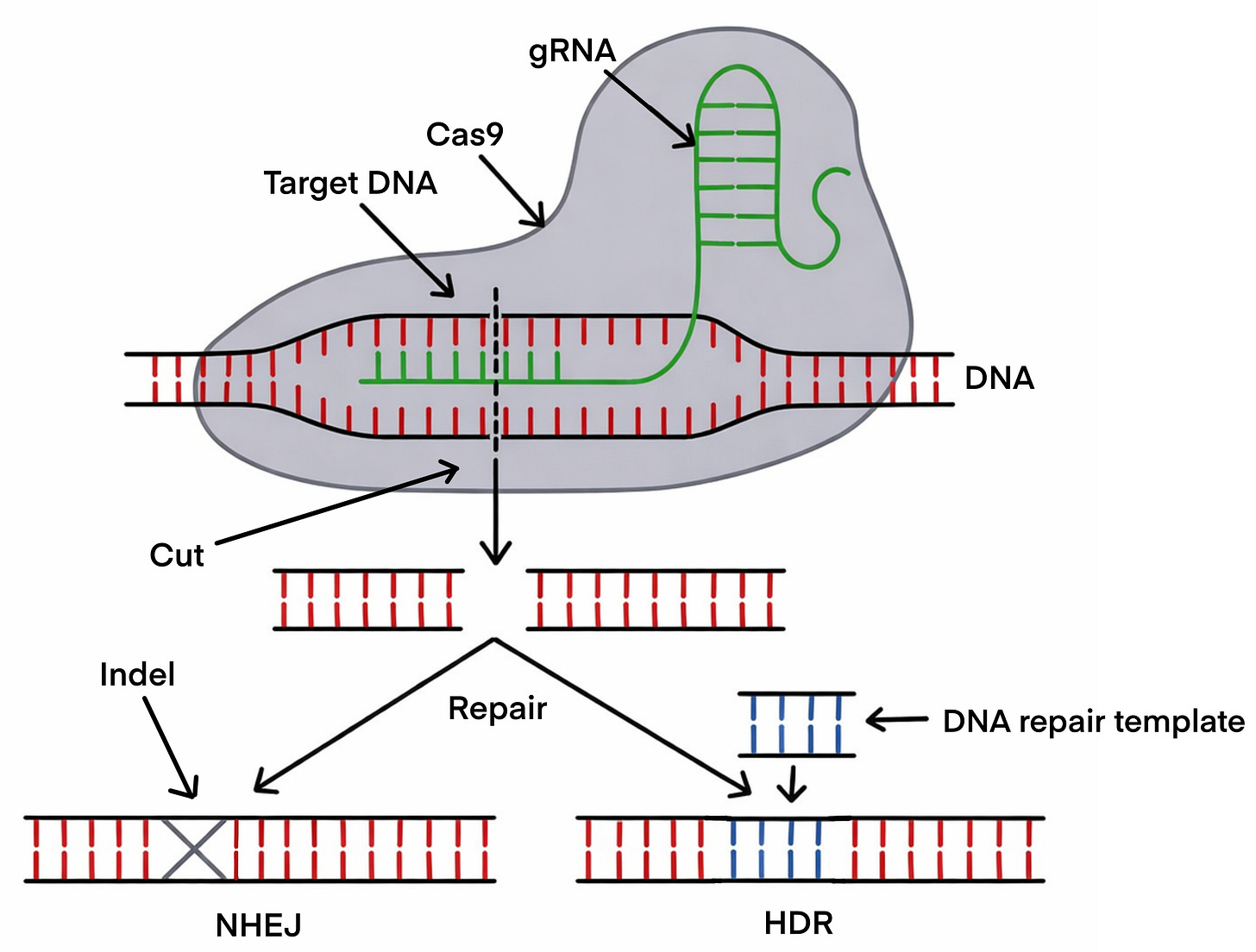
The mechanics are straightforward. You design a guide RNA — about twenty nucleotides long — that matches the genomic sequence you want to target. You introduce the guide RNA and the Cas9 protein into a cell. The complex scans the genome, finds its target, binds to it, and Cas9 cuts through both strands of the DNA double helix at that exact location.
Then the cell notices it has a broken chromosome. And it tries to fix it.
Two Ways to Repair a Break — Two Different Outcomes
What happens next depends on which repair pathway the cell uses, and this is where the actual editing occurs.
The first pathway is non-homologous end joining (NHEJ): the cell’s emergency response, fast and imprecise. It glues the broken ends back together, but the repair is often slightly imperfect — small insertions or deletions appear at the junction. If the cut falls within a gene’s coding sequence, these errors typically knock the gene out entirely. NHEJ is the tool of choice when the goal is disruption: silencing a gene, disabling a regulatory element, removing a repressor.
The second pathway is homology-directed repair (HDR): slower, more demanding, but precise. If you provide a DNA template alongside the CRISPR machinery, the cell can use it as a blueprint to repair the break — incorporating your desired sequence change in the process. HDR can correct a single mutated nucleotide back to its normal form. It is, however, less efficient than NHEJ and works best in dividing cells.
Think of it this way. NHEJ is like taping a torn page back together — fast, functional, but the join is never quite clean. HDR is like locating the original manuscript, cutting out the damaged line, and replacing it with a perfect copy.
Why Stem Cells Are the Perfect Partner
Genome editing can be performed in virtually any cell type. But stem cells — particularly induced pluripotent stem cells (iPSCs) and hematopoietic stem cells (HSCs) — have properties that make them uniquely suited as an editing platform.
iPSCs can be expanded indefinitely. That matters enormously. After introducing a CRISPR edit, researchers can grow thousands of individual cell clones, screen each one to confirm the edit is correct, check the rest of the genome for unintended off-target changes, and select only the verified clones to take forward. In a primary cell with a limited lifespan in culture, that level of quality control would be impossible.
Once you have a well-characterized, correctly edited iPSC clone, its pluripotency does the rest: differentiate it into neurons, heart muscle cells, pancreatic beta cells — whatever cell type the disease affects. The edited genome is present in every daughter cell.
For blood disorders, HSCs are the natural target for a different reason: correct the mutation in HSCs and transplant them back into a patient, and the corrected cells repopulate the entire blood and immune system. It is, in a sense, a permanent fix delivered through the body’s own biology.
That principle is exactly what the first approved CRISPR therapy was built on.
From Laboratory to Clinic: The First Approval
In December 2023, the FDA approved Casgevy (exa-cel) — a CRISPR-based therapy for sickle cell disease and transfusion-dependent beta-thalassemia, developed by Vertex Pharmaceuticals and CRISPR Therapeutics. It was the first CRISPR medicine approved anywhere in the world, and its approval came faster than almost anyone in the field had expected.
Both diseases are caused by mutations in hemoglobin — the protein that carries oxygen in red blood cells. In sickle cell disease, a single nucleotide change causes hemoglobin to polymerize under low oxygen conditions, contorting red blood cells into rigid, crescent-shaped structures that block blood vessels, cause excruciating pain crises, and damage organs over decades. Beta-thalassemia results in insufficient hemoglobin production, leaving patients dependent on regular blood transfusions throughout their lives.
The therapeutic strategy behind Casgevy is conceptually elegant. Rather than trying to correct the mutated adult hemoglobin gene directly, it reactivates a gene the body already has — but switched off.
Fetuses produce a form of hemoglobin — fetal hemoglobin — that works beautifully and is unaffected by the mutations that cause sickle cell disease or beta-thalassemia. After birth, a transcriptional repressor called BCL11A silences the fetal hemoglobin genes, and adult hemoglobin takes over. Casgevy uses CRISPR to disrupt BCL11A in a patient’s own HSCs — reawakening fetal hemoglobin production and compensating for the defective adult form.
The process: harvest HSCs from the patient, edit them outside the body in a specialized manufacturing facility, administer conditioning chemotherapy to make space in the bone marrow, and reinfuse the edited cells. Early clinical data showed that the vast majority of treated patients became transfusion-independent or pain crisis-free — outcomes that represent a profound change in quality of life for people who have often spent their entire lives managing one of the most debilitating inherited diseases in existence.
Beyond Cutting: Sharper Tools
CRISPR-Cas9 in its original form is a blunt instrument. It cuts both strands of DNA and relies on the cell’s repair machinery to introduce the desired change — a process that is powerful but imprecise, and carries some risk of unintended consequences.
The field has not stood still.
Base editing, developed primarily by David Liu’s laboratory at the Broad Institute, uses a modified Cas9 that does not cut DNA. Instead, it chemically converts one nucleotide to another — C to T, or A to G — at a specific location, without ever creating a double-strand break. Many disease-causing mutations are single nucleotide changes, and base editing can correct them with greater precision and fewer unintended effects than standard CRISPR cutting.
Prime editing goes further still. Also from Liu’s laboratory, it fuses a modified Cas9 to a reverse transcriptase enzyme, guided by an extended RNA template encoding the desired edit. Prime editing can introduce virtually any small change — substitutions, small insertions, small deletions — at a defined location, again without a double-strand break. It has been described, fairly, as a “search and replace” function for the genome.
Both technologies are now in active clinical development. Prime editing, in particular, has just reached a significant milestone — one I’ll cover in detail in a dedicated post, because it deserves its own space.
The Line That Should Not Have Been Crossed
Not all uses of CRISPR have been sanctioned. In November 2018, a Chinese scientist named He Jiankui announced at an international conference in Hong Kong that he had edited the genomes of human embryos — embryos that were subsequently implanted and brought to term. Two girls, referred to in press reports as Lulu and Nana, were born with edited genomes. A third edited pregnancy followed.
The target was the CCR5 gene, which encodes a co-receptor that HIV uses to enter cells. He’s stated goal was to confer HIV resistance.
The response from the international scientific community was immediate, nearly unanimous, and overwhelmingly negative.
The reason this crossed a fundamental line is not technical — it is categorical. Editing an embryo means editing the germline: changes present in every cell of the resulting human, including their reproductive cells, heritable by their children and every generation that follows. This is categorically different from the somatic editing used in therapies like Casgevy, which affects only the treated patient and goes no further.
Germline editing of this kind was — and remains — prohibited in most countries and explicitly condemned by major scientific and bioethics bodies. Beyond the regulatory violation, the scientific rationale was deeply questionable: effective HIV prevention already exists, the editing was imprecise by the standards of the time, and the risks to a developing embryo with no say in the matter were not justified by any genuine medical necessity.
He Jiankui was convicted of illegal medical practice by a Chinese court and sentenced to three years in prison.
The episode is a cautionary tale that the scientific community returns to often. CRISPR makes germline editing technically feasible. Feasibility is not justification. The boundary between somatic and germline editing is not merely a regulatory line — it reflects a deeper ethical commitment to the difference between treating a patient and altering a lineage.
Where We Are
In less than fifteen years, CRISPR-Cas9 has moved from a description of a bacterial immune mechanism to an approved medicine transforming the lives of patients with some of the most severe genetic diseases in existence. The speed of that translation is historically unusual, and it reflects both the power of the underlying biology and the fortunate convergence with iPSC technology — which provided a platform for editing, screening, and manufacturing at the scale medicine requires.
The tools are becoming more precise. Base editing and prime editing are moving through the clinic. The list of diseases in active CRISPR programs continues to expand.
What is striking — and worth holding onto — is that all of this traces back to bacteria defending themselves against viruses. Biology has been running this molecular search-and-destroy system for billions of years. We simply learned to read the manual
Background and further reading
Jinek M, et al. “A programmable dual-RNA–guided DNA endonuclease in adaptive bacterial immunity.” Science. 2012.
Cong L, et al. “Multiplex genome engineering using CRISPR/Cas systems.” Science. 2013.
Doudna JA & Charpentier E. “The new frontier of genome engineering with CRISPR-Cas9.” Science. 2014.
Frangoul H, et al. “CRISPR-Cas9 gene editing for sickle cell disease and β-thalassemia.” New England Journal of Medicine. 2021.
Komor AC, et al. “Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage.” Nature. 2016.
Anzalone AV, et al. “Search-and-replace genome editing without double-strand breaks or donor DNA.” Nature. 2019.
Lander ES, et al. “Adopt a moratorium on heritable genome editing.” Nature. 2019.
Cyranoski D & Ledford H. “Genome-edited baby claim provokes international outcry.” Nature. 2018.
