The Genes That Forgot What They Were For
Humans cannot make their own vitamin C. Nearly every other mammal on Earth can. The reason why is written into our DNA — in the form of a gene that still exists, but no longer works.
A Genome Full of Ghosts
When scientists sequenced the human genome in the early 2000s, they expected to find genes — the functional instructions that encode proteins and drive the machinery of life. What they did not fully anticipate was the sheer number of broken genes they would find alongside the working ones.
The human genome contains hundreds of pseudogenes: DNA sequences that bear the unmistakable hallmarks of once-functional genes — the right structure, the right location, the right evolutionary context — but that have accumulated mutations rendering them silent. They are molecular fossils, preserved in our chromosomes long after the functions they once encoded became unnecessary, or were simply lost by chance.
They are, in a very real sense, the genomic record of our evolutionary past.
The Vitamin C Gene We No Longer Use
The story of vitamin C is one of the most instructive examples in all of evolutionary biology.
Most mammals synthesize their own ascorbic acid — vitamin C — in the liver. The final step in this biosynthetic pathway is catalyzed by an enzyme called L-gulonolactone oxidase, encoded by the GULO gene. Rats make it. Mice make it. Dogs, cows, and horses make it. They never need to obtain vitamin C from their diet, because their cells manufacture it continuously.
Humans cannot do this. Neither can other great apes, nor guinea pigs, nor certain bat species. We are entirely dependent on dietary sources for a molecule that is essential for collagen synthesis, immune function, and a dozen other critical biological processes. A deficiency kills — historically, through the disease scurvy, which devastated sailors on long voyages before the role of citrus fruits was understood.
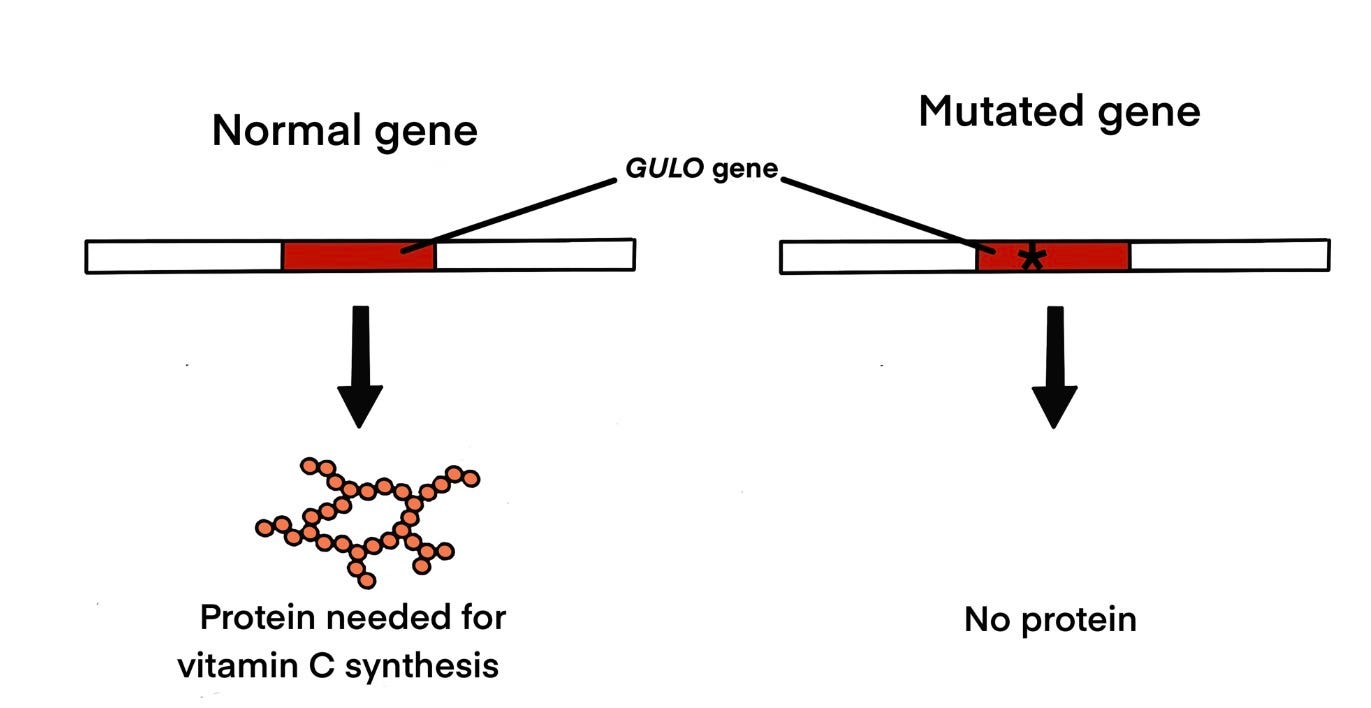
Why can’t we make our own? Not because we lack the gene. We have it — or rather, we have the ruins of it. The human genome contains a clear remnant of the GULO gene at chromosome 8, in exactly the position one would predict based on synteny with other mammals. But a series of mutations — including deletions, stop codons, and frameshift mutations — have disabled it completely. The gene is there. It just doesn’t work.
This is not unique to humans. When researchers examined the broken GULO gene across primates and guinea pigs, they found something remarkable: the same mutations are shared across species that share a common ancestry. The great apes carry overlapping sets of the same deletions. This is precisely what one would predict if a single mutational event — or a cascade of events — disabled the gene in a common ancestor, and that broken version was then inherited by all descendants. It is the molecular equivalent of finding the same typo in every copy of a book — evidence of a shared origin, not independent invention.
Not One Broken Gene, But Hundreds
The GULO pseudogene is striking because its functional consequences are so visible — we take vitamin supplements where our ancestors needed none. But it is far from unique.
The human genome contains somewhere in the range of 10,000–20,000 pseudogenes, depending on how they are counted and classified. Many are processed pseudogenes — copies of genes that were reverse-transcribed from messenger RNA and reinserted into the genome, stripped of the regulatory sequences needed to function. Others, like GULO, are unprocessed pseudogenes: the original gene, sitting in its original chromosomal location, progressively dismantled by mutation over evolutionary time.
Among the most illuminating examples are our olfactory receptor genes — the genes that encode the proteins used to detect smell.
The Nose That Evolution Walked Away From
Smell is mediated by a large family of receptor proteins embedded in the membranes of olfactory neurons in the nasal epithelium. Each receptor is encoded by a separate gene. The more functional olfactory receptor genes a species has, the richer and more discriminating its sense of smell.
Mice have approximately 1000 functional olfactory receptor genes. They live in a world saturated with chemical information — detecting predators, locating food, and navigating social hierarchies largely through smell. Rats are similar. So are most other non-primate mammals.
Humans have roughly 400 functional olfactory receptor genes — until you look at what sits alongside them in the genome. We carry a similar number of olfactory receptor pseudogenes: the broken remains of genes that were fully functional in our ancestors, gradually inactivated as primate evolution shifted its investment from smell toward vision and the expansion of visual cortex. The transition to trichromatic color vision in primates — the ability to distinguish red, green, and blue — appears to have coincided with a relaxation of selective pressure on olfaction. If you can see ripe fruit, you need your nose less.
The pseudogenes are still there, though. They sit in the same genomic clusters as the functional receptors, recognizable by their sequence as olfactory genes, but silenced by the same kinds of mutations that disabled GULO: premature stop codons, frameshifts, deletions in critical exons. They are the olfactory architecture of an ancestor that lived in a world where smell mattered more.
Why Pseudogenes Matter for Understanding Evolution
Pseudogenes are sometimes dismissed as “junk DNA” — genomic noise with no biological significance. This framing is increasingly outdated. Some pseudogenes appear to have acquired regulatory functions, producing non-coding RNAs that influence the expression of neighboring genes. The biology is more complex than a simple on/off switch.
But their primary significance remains evolutionary, and it is profound.
A pseudogene is a prediction. If two species share a common ancestor, and that ancestor had a functional gene, then both species should carry recognizable remnants of that gene in the same chromosomal location — even if it has been degraded by different mutations in each lineage. And if a pseudogene arose by mutation in that common ancestor, both descendant species should carry the same core mutations, with additional, species-specific degradation layered on top.
This is exactly what we observe. Again and again, across genes and across species, the pattern holds. Pseudogenes are not random noise. They are a nested record of shared ancestry, written into the genome in a form that is, if anything, more informative than functional genes — because unlike functional genes, pseudogenes are free to accumulate mutations without constraint. They degrade at a predictable rate, tracking evolutionary time.
The broken GULO gene in your chromosome 8 is not a flaw in an otherwise perfect design. It is a historical document — a record of a mutation that occurred in a primate ancestor tens of millions of years ago, and has been faithfully copied into every descendant genome since.
Including yours.
What the Silence Tells Us
There is something genuinely remarkable about pseudogenes: they are useful precisely because they have stopped being useful. A gene under strong selective pressure stays functional because mutations that break it are eliminated from the population. A pseudogene has no such constraint. It accumulates mutations neutrally, freely, and in proportion to elapsed evolutionary time.
This makes pseudogenes one of the most powerful molecular tools for reconstructing evolutionary history. They let us date divergence events, trace migrations, and compare lineages with a resolution that functional genes — conserved and maintained by selection — cannot always provide.
But perhaps their deepest message is simpler than any of that. The human genome is not an optimized blueprint, engineered from scratch. It is a heavily annotated palimpsest: layer upon layer of evolutionary history, with old instructions preserved even when they are no longer followed. The ghosts of ancestral capabilities — to make vitamin C, to smell the world in high resolution, and hundreds of other things we have lost — are all still there, written in the sequence.
The genome remembers what evolution forgot.
Background and further reading
Nishikimi, M., Fukuyama, R., Minoshima, S., Shimizu, N., & Yagi, K. Cloning and chromosomal mapping of the human nonfunctional gene for L-gulonolactone oxidase, the enzyme for L-ascorbic acid biosynthesis missing in man. Journal of Biological Chemistry, 1994.
Gilad, Y., Wiebe, V., Przeworski, M., Lancet, D., & Pääbo, S. Loss of olfactory receptor genes coincides with the acquisition of full trichromatic vision in primates. PLOS Biology, 2004.
Torrents, D., Suyama, M., Zdobnov, E., & Bork, P. A genome-wide survey of human pseudogenes. Genome Research, 2003.
Zhang, Z., & Gerstein, M. Large-scale analysis of pseudogenes in the human genome. Current Opinion in Genetics & Development, 2004.
