The Genome Is Not Yours Alone
Nearly half the DNA in your cells did not originate in any human ancestor. It arrived from elsewhere — and some of it is still doing a job.
A Patchwork Inheritance
When scientists completed the sequencing of the human genome in the early 2000s, one of the most startling findings had nothing to do with our protein-coding genes. It was the sheer volume of foreign-looking sequence — DNA whose structure, organization, and evolutionary history pointed not to any human or primate ancestor, but to viruses that had infected our ancestors’ germlines across hundreds of millions of years.
The human genome is, in a very real sense, a patchwork. It carries not just the instructions for making a human being, but the molecular residue of countless infections that left their mark on the germ cells of our ancestors. To read the human genome is to read a layered archaeological record — part human, part viral, part history of the planet itself.
Ancient Passengers
Start with the most dramatic number: approximately 8% of the human genome consists of sequences derived from ancient retroviruses. That is more than twice the amount of DNA devoted to protein-coding genes.
Retroviruses replicate by a distinctive mechanism. After infecting a cell, they convert their RNA genome into DNA using an enzyme called reverse transcriptase, then insert that DNA into the host cell’s chromosomes. If a retrovirus manages to infect a germ cell — a sperm or egg cell — the integrated viral DNA is inherited by every cell in the offspring’s body, and potentially by every generation that follows. It becomes a permanent passenger.
These integrated sequences are called human endogenous retroviruses, or HERVs. Over tens of millions of years, they accumulated in our lineage — integrating, duplicating, mutating. The vast majority are now badly degraded: stripped of functional genes, riddled with mutations, recognizable as viral only by their characteristic structural features. They are genomic fossils.
When you add the broader category of all transposable elements — mobile DNA sequences including LINE elements and Alu repeats — the total rises to nearly 45% of the human genome. Close to half our DNA is, at its origin, mobile or parasitic sequence that has been accumulating since before our vertebrate ancestors emerged.
The real significance of these sequences is not their abundance — it is their location. Where exactly a retrovirus integrated, and which species share that exact location, turns out to be one of the most powerful tools in evolutionary biology.
A Molecular Phylogenetic Tree, Written in Viral Scars
When a retrovirus integrates into a germ cell, it inserts at a specific, essentially random location in a genome of three billion base pairs. The probability of two independent integration events landing at precisely the same position in a genome of three billion base pairs is vanishingly small — effectively zero for practical purposes. If two species share an endogenous retroviral insertion at the same genomic location, the only parsimonious explanation is that the integration occurred once, in a common ancestor, and was then inherited by both lineages.
This is exactly the pattern we find. And the distribution of shared insertions maps onto the tree of life with remarkable precision.
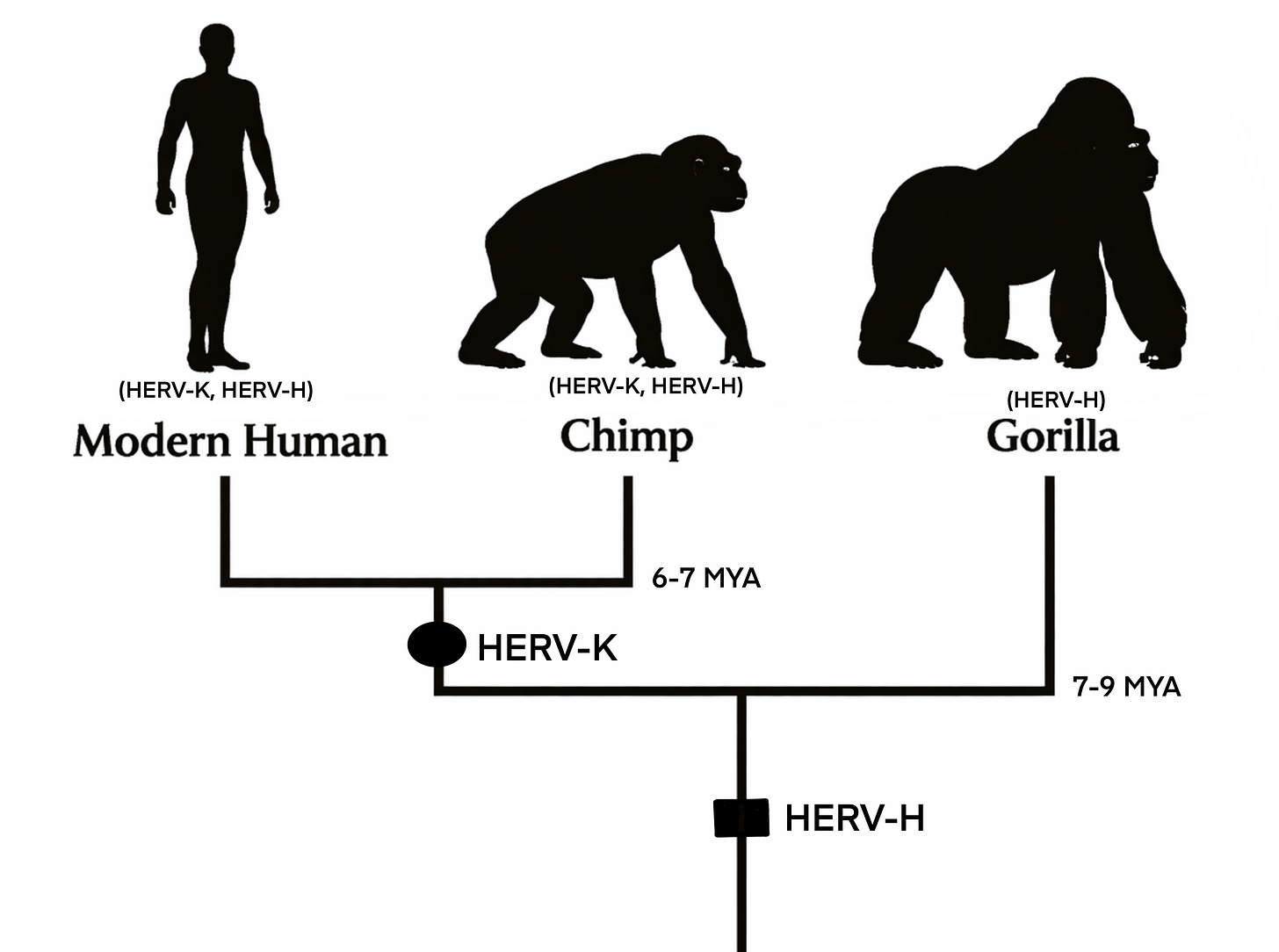
Some HERV insertions are shared across humans, chimpanzees, and gorillas — integrated millions of years before these lineages diverged. HERV-H is one example, found at equivalent positions in all three species. Others, like HERV-K(HML-2), are shared only between humans and chimpanzees, reflecting integrations that occurred after the gorilla lineage branched off roughly 7–9 million years ago but before humans and chimpanzees split around 6–7 million years ago (Figure below). A subset of insertions are found in humans alone — integrations that occurred after our lineage diverged from chimpanzees.
Move further back, and some HERVs are shared across all great apes, others across all primates including Old World monkeys, and some extend into other placental mammals entirely — marking infections in ancestors that lived tens of millions of years before any of the modern lineages existed.
The result is a perfect phylogenetic tree — not reconstructed from protein sequences or anatomical features, but read directly from the chromosomal addresses of ancient viral integrations. Each shared insertion is a timestamp; together they define a branching hierarchy entirely consistent with every other line of evolutionary evidence, and explicable only by common descent.
This is not an argument from similarity. It is an argument from shared, positionally identical molecular events. A random retroviral insertion has no function to converge on — it is simply a mark, left at a moment in time, inherited by every descendant since.
When a Virus Becomes a Gene
Not all of this inherited viral DNA is inert. Some has been repurposed — captured by evolution and put to work in ways that have nothing to do with the virus’s original agenda.
The most remarkable example involves the placenta.
The placenta requires a tissue layer called the syncytiotrophoblast: a continuous, multinucleated sheet of cells that forms the interface between fetal and maternal bloodstreams. For this layer to form, individual trophoblast cells must fuse together — a process of controlled cell fusion that few biological structures require.
The protein that drives this fusion in humans is called syncytin-1. It is encoded by a gene derived from the env gene of an ancient HERV — specifically HERV-W, which integrated into the primate germline approximately 25 million years ago. The env gene originally mediated viral entry into host cells by fusing the viral envelope with the cell membrane. Evolution found a new use for this membrane-fusion machinery: building the placenta.
What makes this especially remarkable is that it happened more than once. Mice and rats have their own syncytins — encoded by a completely different set of endogenous retroviruses that integrated independently into the rodent lineage roughly 20 million years ago. Rabbits have theirs. Ruminants have yet another. Unrelated retroviruses, integrated into unrelated mammalian lineages at unrelated times, each independently co-opted for the same biological function through convergent evolution. When syncytin genes are knocked out in mice, the animals die in utero because their placentas fail to form.
A viral gene, captured from an ancient infection, is now essential for mammalian reproduction. Without it, you would not exist.
The Genome Remembers
There is no clean boundary between “our” DNA and “their” DNA. The genome is a palimpsest — a document written, overwritten, and annotated across geological time by many hands, most of them viral.
Some of this inherited sequence is genuinely inert: molecular debris that has been accumulating since before our vertebrate ancestors emerged. But some of it has been repurposed, domesticated, folded into the machinery of life itself. The gene that builds your placenta was once a viral infection. The timestamps written into your chromosomal architecture reconstruct the entire branching history of the primate lineage, one integration event at a time.
This is what makes genomics genuinely humbling. The human genome is not a blueprint engineered from scratch. It is an archive — of ancient infections, evolutionary accidents, and opportunistic co-options that happened to persist long enough to become part of what we are.
The viral scars are not blemishes on an otherwise clean design. In many cases, they are the design.
Background and further reading
Mi, S., Lee, X., Li, X., Veldman, G.M., Finnerty, H., Racie, L., LaVallie, E., Tang, X.Y., Edouard, P., Howes, S., Keith, J.C., & McCoy, J.M. Syncytin is a captive retroviral envelope protein involved in human placental morphogenesis. Nature, 2000.
Feschotte, C., & Gilbert, C. Endogenous viruses: insights into viral evolution and impact on host biology. Nature Reviews Genetics, 2012.
Lander, E.S., et al. Initial sequencing and analysis of the human genome. Nature, 2001.
Johnson, W.E., & Coffin, J.M. Constructing primate phylogenies from ancient retrovirus sequences. Proceedings of the National Academy of Sciences, 1999.
Dupressoir, A., Marceau, G., Vernochet, C., Bénit, L., Kanellopoulos, C., Sapin, V., & Heidmann, T. Syncytin-A and syncytin-B, two fusogenic placenta-specific murine envelope genes of retroviral origin conserved in Muridae. Proceedings of the National Academy of Sciences, 2005.
Lebedev, Y.B., Belonovitch, O.S., Zybrova, N.V., Khil, P.P., Kurdyukov, S.G., Vinogradova, T.V., Hunsmann, G., & Sverdlov, E.D. Differences in HERV-K LTR insertions in orthologous loci of humans and great apes. Gene, 2000.
